Random Forest in R – Understand every aspect related to it!
We offer you a brighter future with FREE online courses - Start Now!!
We will study the concept of random forest in R thoroughly and understand the technique of ensemble learning and ensemble models in R Programming. We will also explore random forest classifier and process to develop random forest in R Language.
So, let’s start.
Introduction to Random Forest in R
What are Random Forests?
The idea behind this technique is to decorrelate the several trees. Ensemble technique called Bagging is like random forests. It is generated on the different bootstrapped samples from training data. And, then we reduce the variance in trees by averaging them. Hence, in this approach, it creates a large number of decision trees in R.
Master the concept of R Decision Trees before proceeding further
We use the R package “randomForest” to create random forests.
What is Ensemble Learning in R?
It is a type of supervised learning technique. The basic idea behind it is to generate many models on a training dataset and then combining their output rules.
We use it to generate lots of models by training on Training Set and combining them at the end. Hence, we can use it to improve the predictive performance of decision trees by reducing the variance in the trees through averaging them. It is called the random forest technique.
What are Ensemble Models in R?
It is a type of model which combine results from different models and is usually better than the result from one of the individual models.
Some of the features of random forests in R Programming are as follows:
- It is the type of model which runs on large databases.
- Random forests allow handling of thousands of input variables without variable deletion.
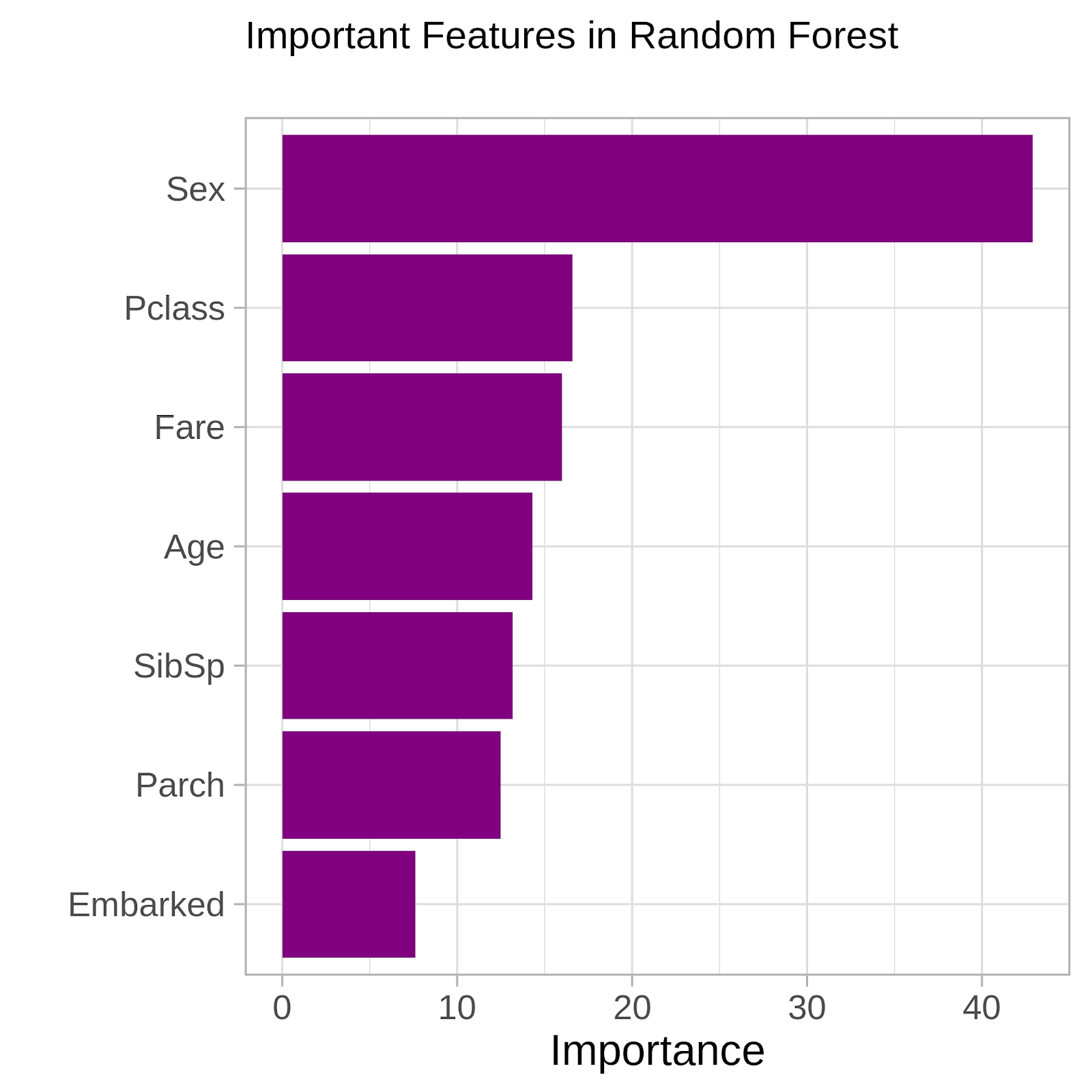
- It gives very good estimates stating which variables are important in the classification.
The tutorial to gain expertise in Classification in R Programming
Random Forest Classifier
At training time, we can classify the ensemble learning method of random forest and thus we can operate it by constructing a multitude of decision trees.
Adele Cutler and Leo Breiman developed it. Here the combination of two different methods is done by Leo’s bagging idea and the random selection of features introduced by Tin Kan Ho. He also proposed Random Decision Forest in the year 1995.
Functions of Random Forest in R
If the number of cases in the training set is N, and the sample N case is at random, each tree will grow. Thus, this sample will be the training set for growing the tree. If there are M input variables, we specify a number m<<M such that at each node, m variables are selected at random out of the M. The value of m is constant during the forest growing and hence, each tree grows to the largest extent possible.
Do you know about Clustering in R Programming Language
Developing Random Forest in R
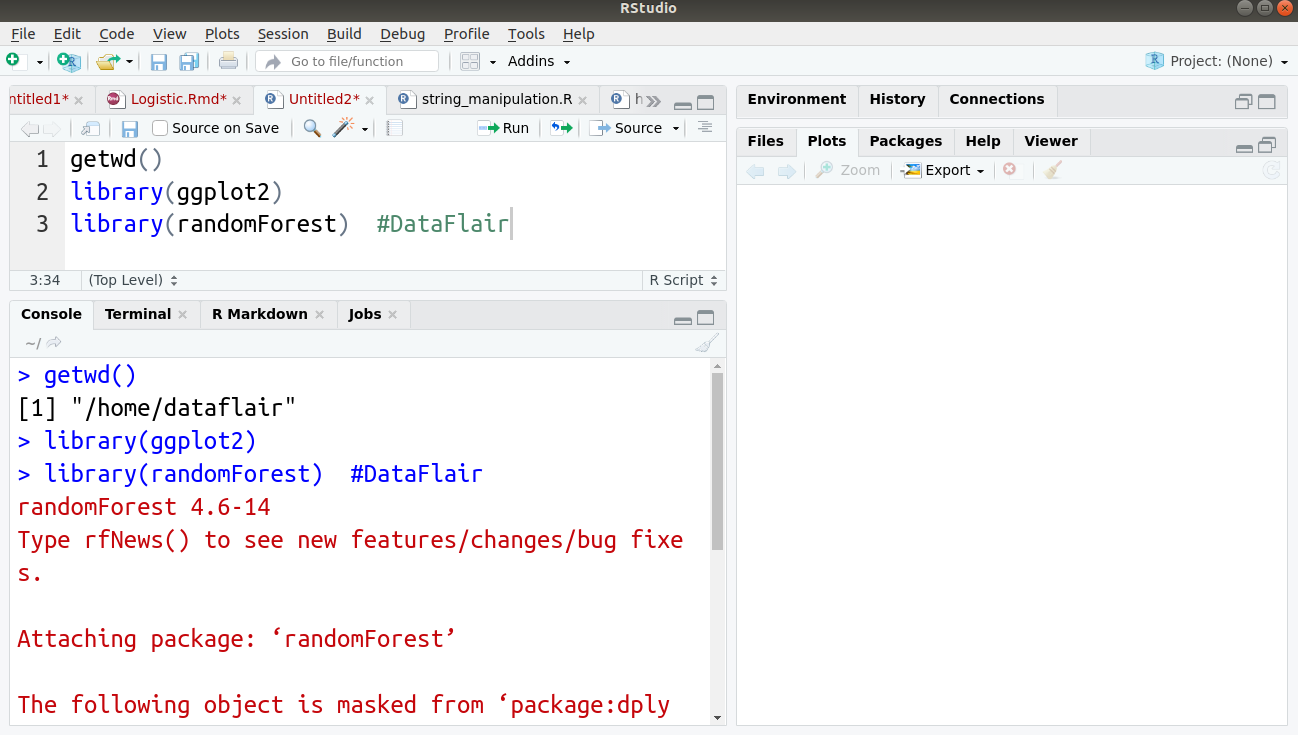
We will first import the important libraries, such as ggplot2 and randomForest as follows:
#Author DataFlair getwd() library(ggplot2) library(randomForest)
Output:
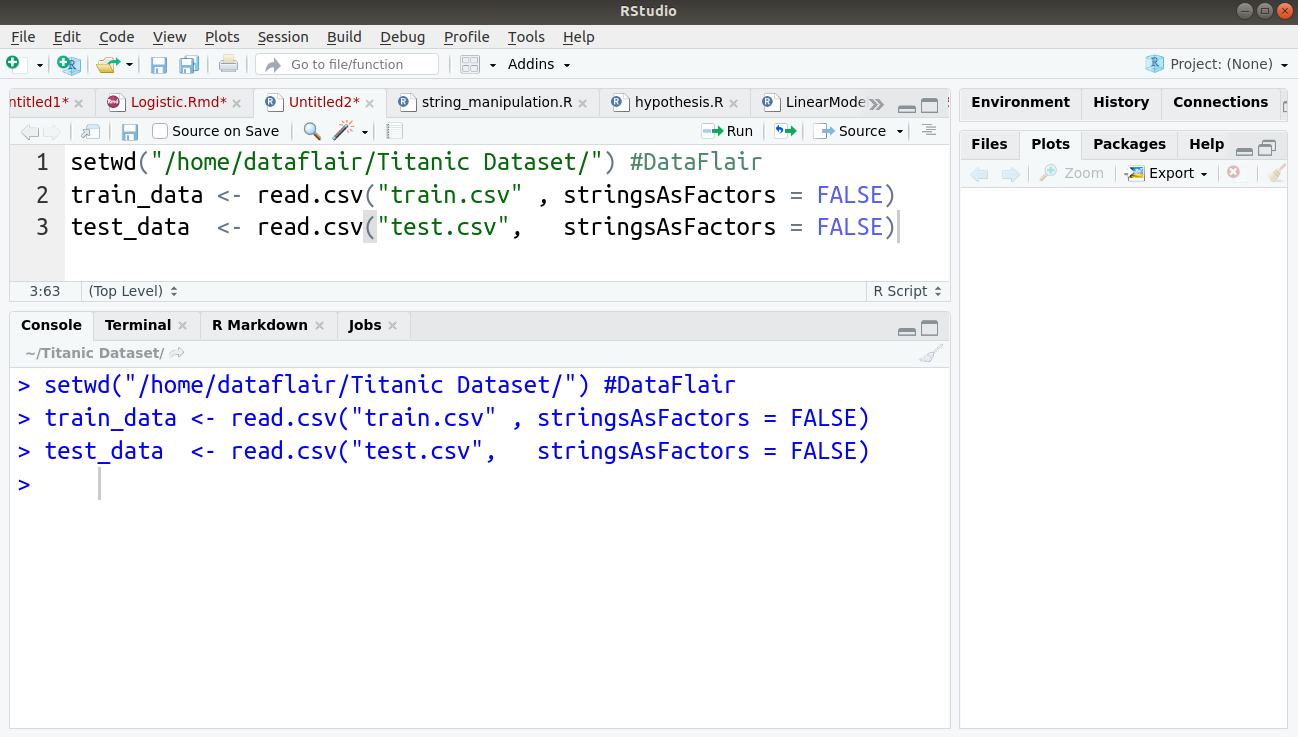
We will use the popular titanic survival prediction dataset and will import the training and test sets into our two corresponding variables:
setwd("/home/dataflair/Titanic Dataset/") #DataFlair
train_data <- read.csv("train.csv" , stringsAsFactors = FALSE)
test_data <- read.csv("test.csv", stringsAsFactors = FALSE)
Output:
You can’t afford to miss the guide on e1071 Package and SVM Training & Testing Models in R
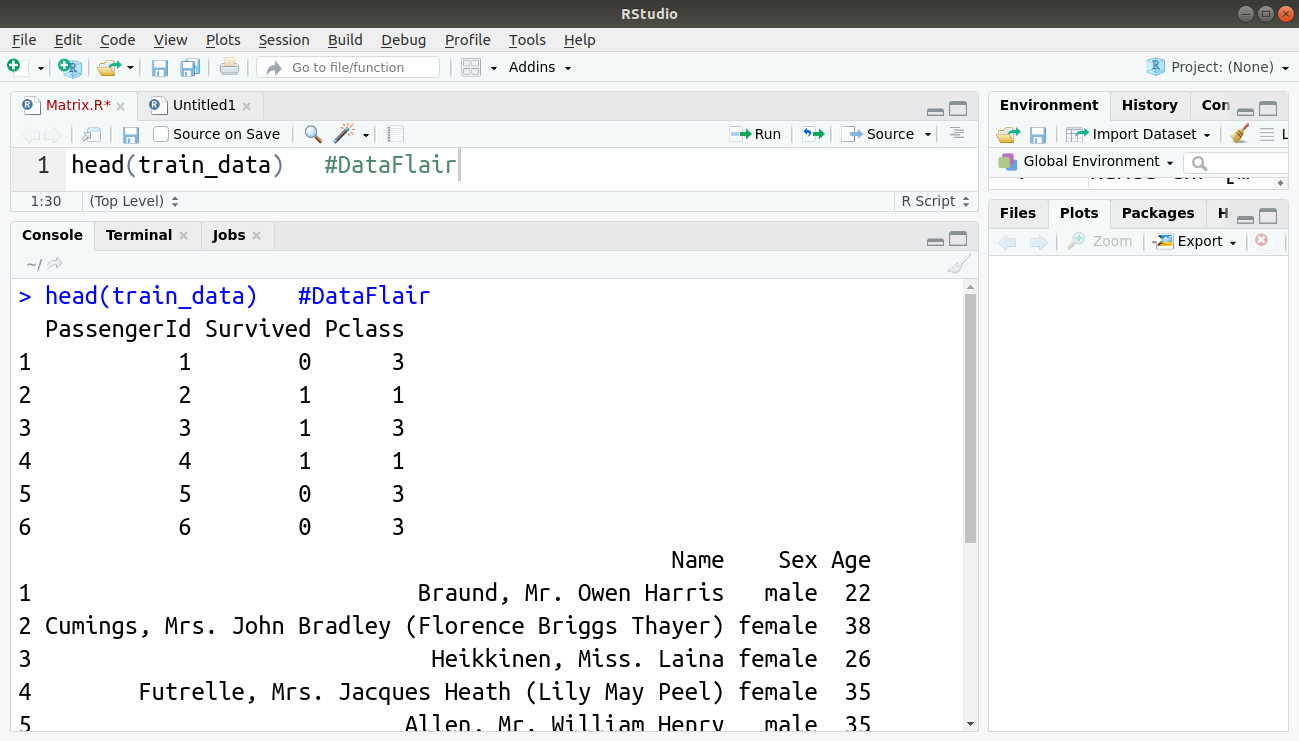
Let us now take a look at the first six entries of our imported training and testing dataset:
> head(train_data)
Output:
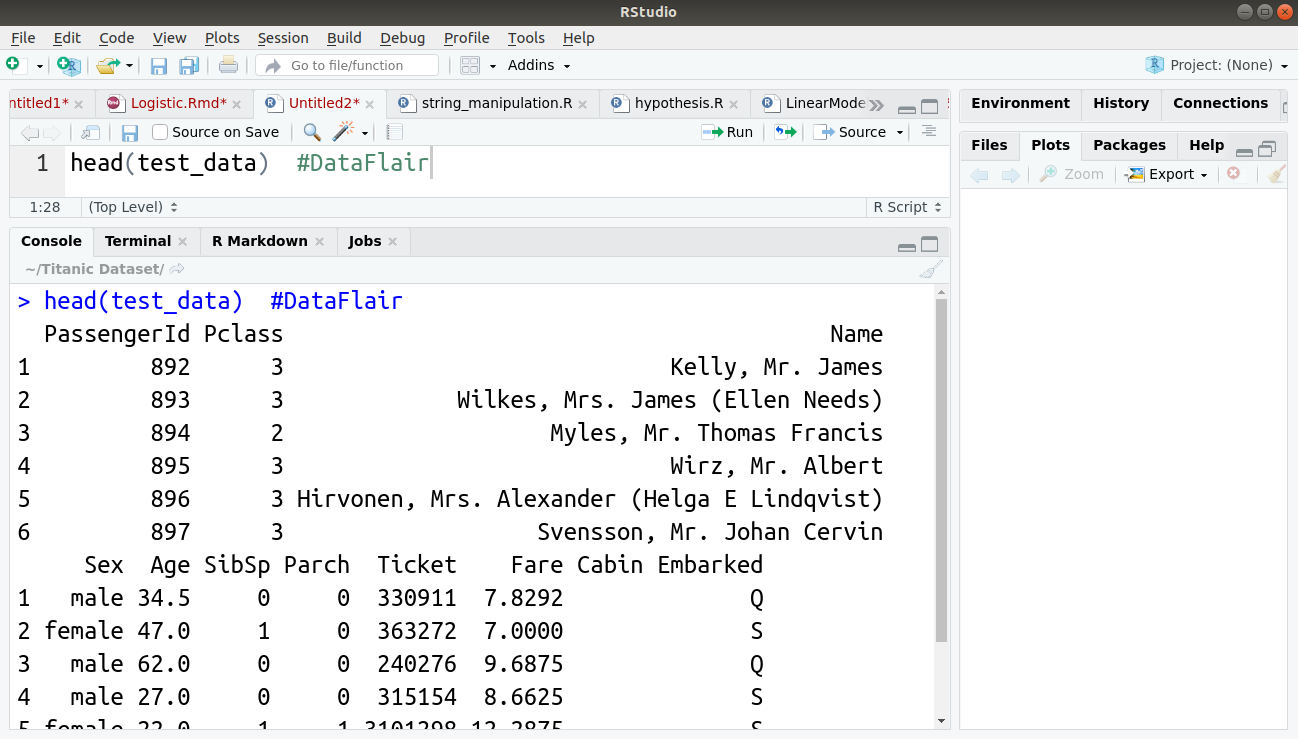
head(test_data)
Output:
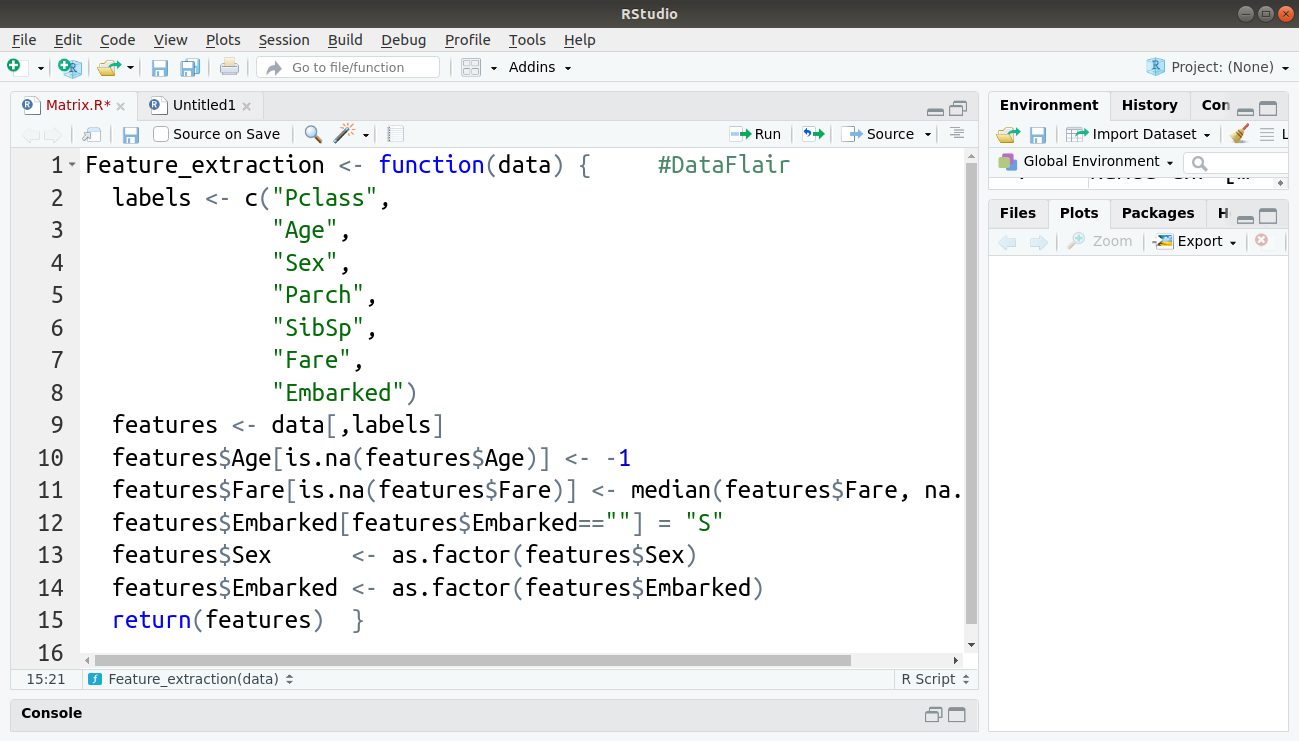
Extraction of features is the next most important step towards building our model. We perform this in the form of a function called ‘Feature_extraction’ that will extract features from whatever input data is provided to it whenever it is called.
Feature_extraction <- function(data) {
labels <- c("Pclass",
"Age",
"Sex",
"Parch",
"SibSp",
"Fare",
"Embarked")
features <- data[,labels]
features$Age[is.na(features$Age)] <- -1
features$Fare[is.na(features$Fare)] <- median(features$Fare, na.rm=TRUE)
features$Embarked[features$Embarked==""] = "S"
features$Sex <- as.factor(features$Sex)
features$Embarked <- as.factor(features$Embarked)
return(features)
}
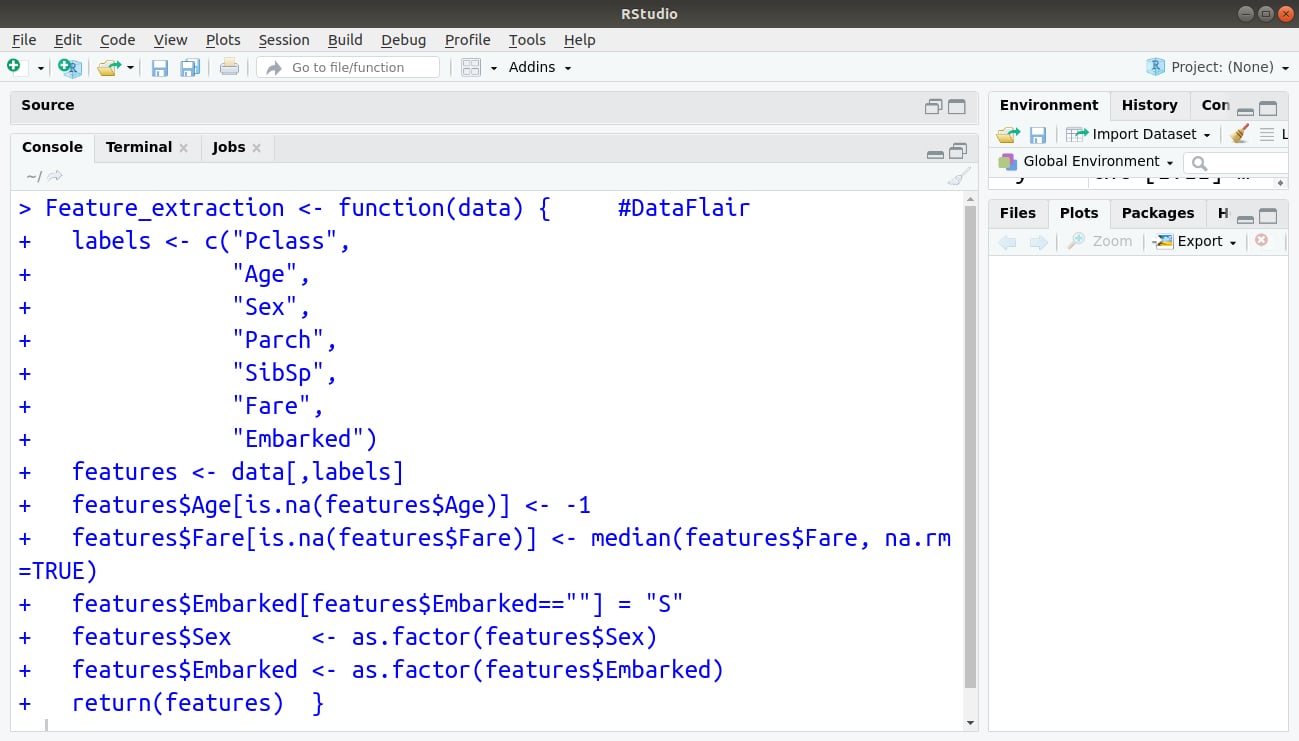
Code Display:
Output:
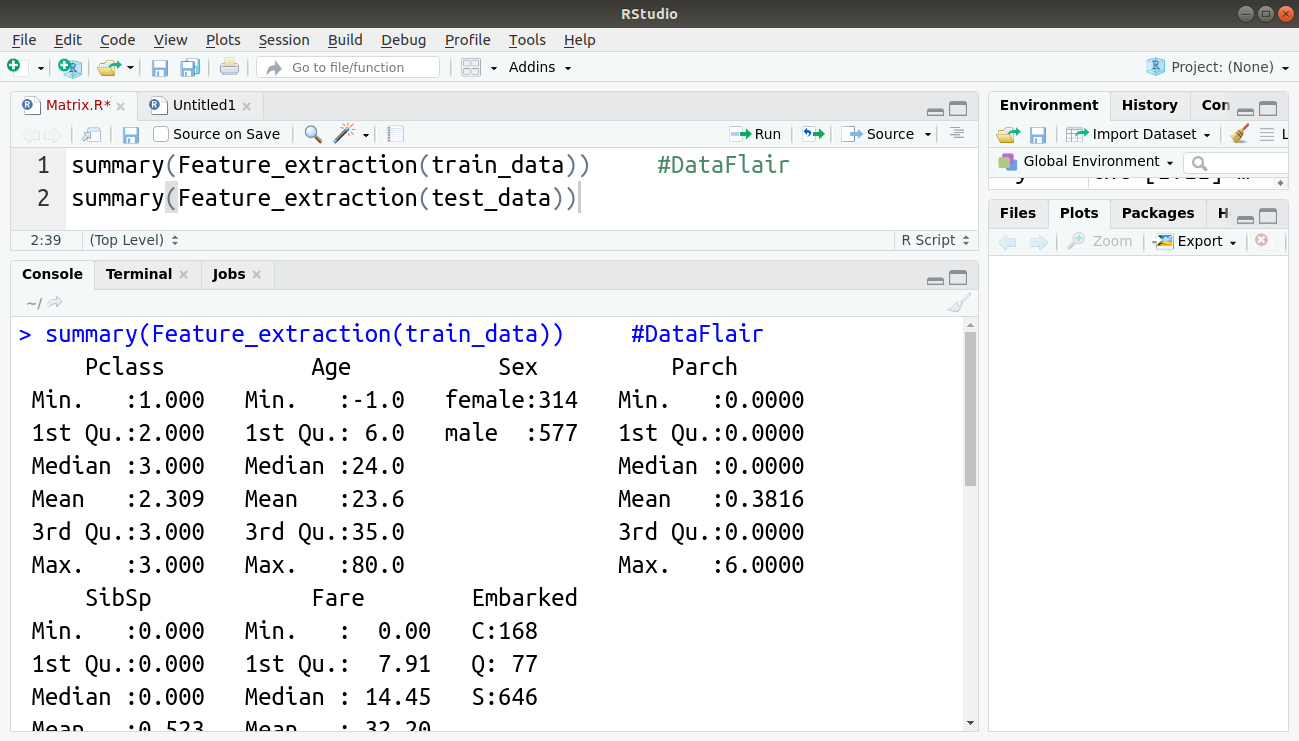
Let us call the above function by displaying the summary of the extracted features of training and test sets:
> summary(Feature_extraction(train_data)) > summary(Feature_extraction(test_data))
Output:
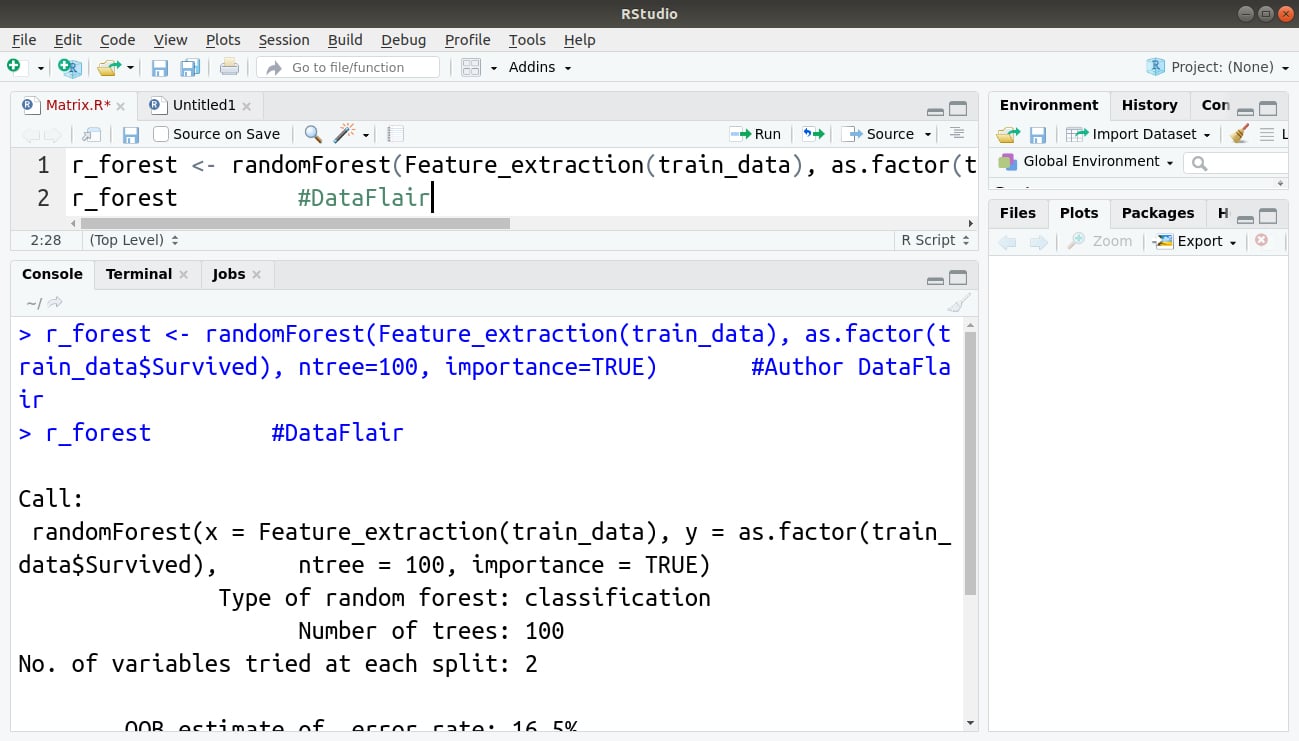
In the next step, we will implement our randomForest() function as follows:
> r_forest <- randomForest(Feature_extraction(train_data), as.factor(train_data$Survived), ntree=100, importance=TRUE) #Author DataFlair > r_forest
Output:
Let us now extract some of the important features with the following lines of code:
> important <- importance(r_forest, type=1 ) #Author DataFlair > Important_Features <- data.frame(Feature = row.names(important), Importance = important[, 1])
Output:
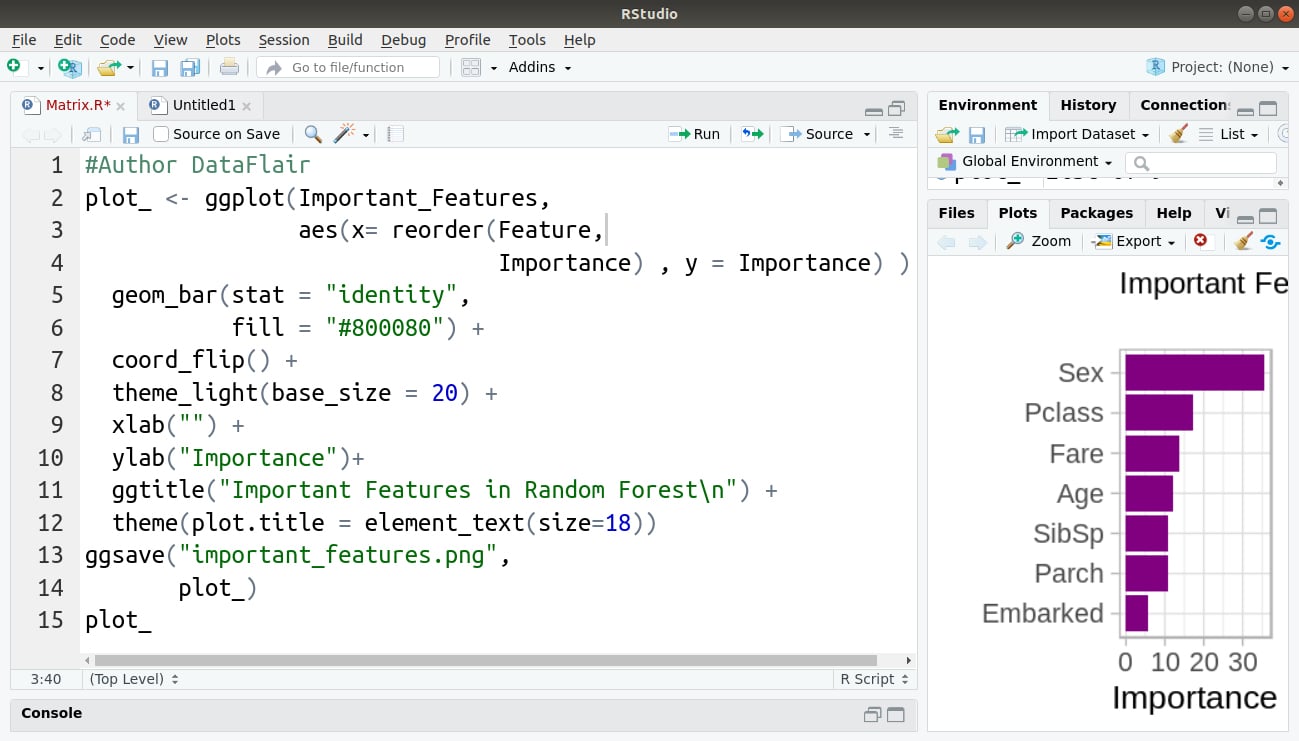
In the final step, we will use ggplot2 to plot our set of important features:
#Author DataFlair
plot_ <- ggplot(Important_Features,
aes(x= reorder(Feature,
Importance) , y = Importance) ) +
geom_bar(stat = "identity",
fill = "#800080") +
coord_flip() +
theme_light(base_size = 20) +
xlab("") +
ylab("Importance")+
ggtitle("Important Features in Random Forest\n") +
theme(plot.title = element_text(size=18))
ggsave("important_features.png",
plot_)
plot_
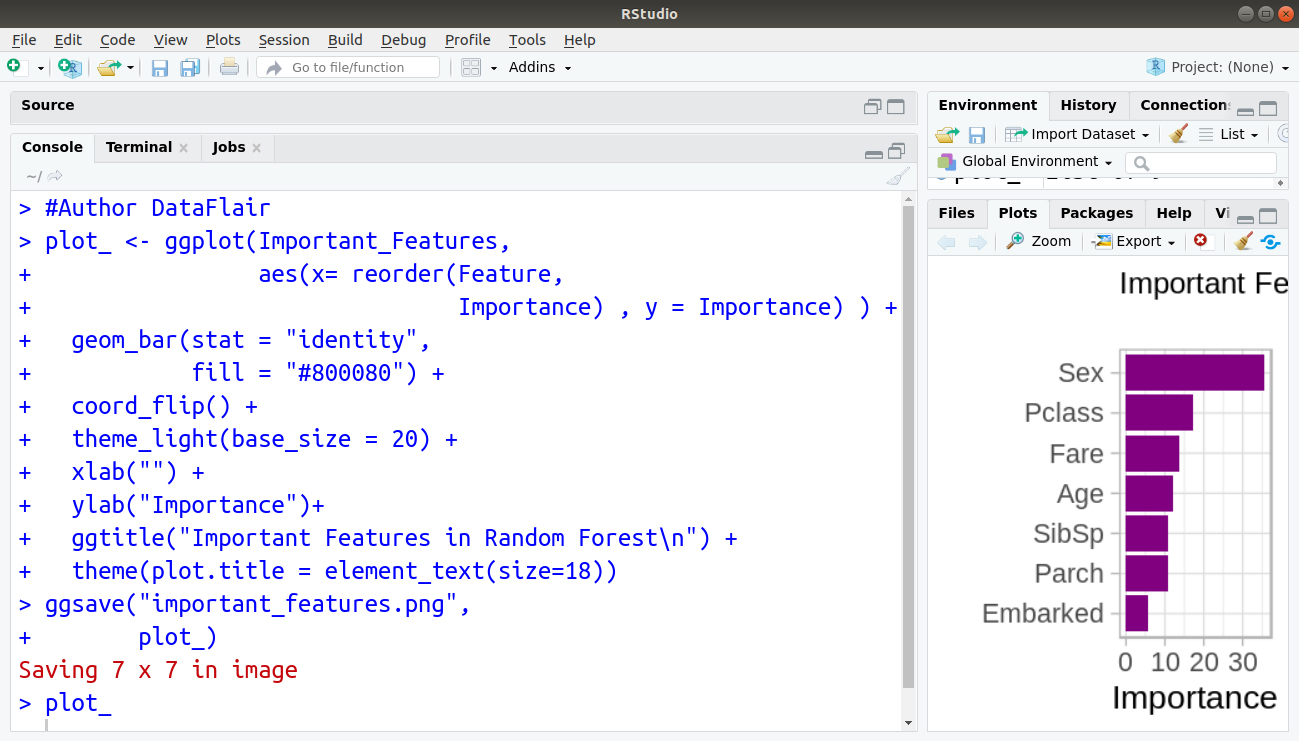
Code Display:
Output:
Bar Graph:
Summary
We have studied the different aspects of random forest in R. We learned about ensemble learning and ensemble models in R Programming along with random forest classifier and process to develop random forest in R.
Now, it’s time to land on Bayesian Network in R
Any queries regarding random forest in R? Enter in the comment section below.
Did you know we work 24x7 to provide you best tutorials
Please encourage us - write a review on Google



where are the data sets in this example?