Project in Python – Breast Cancer Classification with Deep Learning
Machine Learning courses with 100+ Real-time projects Start Now!!
If you want to master Python programming language then you can’t skip projects in Python. After publishing 4 advanced python projects, DataFlair today came with another one that is the Breast Cancer Classification project in Python. To crack your next Python Interview, practice these projects thoroughly and if you face any confusion, do comment, DataFlair is always ready to help you.
Before we begin this Breast Cancer Classification Project in Python, let me provide you the list of advanced python projects published by DataFlair:
- Fake News Detection Python Project
- Parkinson’s Disease Detection Python Project
- Color Detection Python Project
- Speech Emotion Recognition Python Project
- Breast Cancer Classification Python Project
- Age and Gender Detection Python Project
- Handwritten Digit Recognition Python Project
- Chatbot Python Project
- Driver Drowsiness Detection Python Project
- Traffic Signs Recognition Python Project
- Image Caption Generator Python Project
Breast Cancer Classification Project in Python
Get aware with the terms used in Breast Cancer Classification project in Python
What is Deep Learning?
An intensive approach to Machine Learning, Deep Learning is inspired by the workings of the human brain and its biological neural networks. Architectures as deep neural networks, recurrent neural networks, convolutional neural networks, and deep belief networks are made of multiple layers for the data to pass through before finally producing the output. Deep Learning serves to improve AI and make many of its applications possible; it is applied to many such fields of computer vision, speech recognition, natural language processing, audio recognition, and drug design.
What is Keras?
Keras is an open-source neural-network library written in Python. It is a high-level API and can run on top of TensorFlow, CNTK, and Theano. Keras is all about enabling fast experimentation and prototyping while running seamlessly on CPU and GPU. It is user-friendly, modular, and extensible.
Breast Cancer Classification – Objective
To build a breast cancer classifier on an IDC dataset that can accurately classify a histology image as benign or malignant.
Breast Cancer Classification – About the Python Project
In this project in python, we’ll build a classifier to train on 80% of a breast cancer histology image dataset. Of this, we’ll keep 10% of the data for validation. Using Keras, we’ll define a CNN (Convolutional Neural Network), call it CancerNet, and train it on our images. We’ll then derive a confusion matrix to analyze the performance of the model.
IDC is Invasive Ductal Carcinoma; cancer that develops in a milk duct and invades the fibrous or fatty breast tissue outside the duct; it is the most common form of breast cancer forming 80% of all breast cancer diagnoses. And histology is the study of the microscopic structure of tissues.
The Dataset
We’ll use the IDC_regular dataset (the breast cancer histology image dataset) from Kaggle. This dataset holds 2,77,524 patches of size 50×50 extracted from 162 whole mount slide images of breast cancer specimens scanned at 40x. Of these, 1,98,738 test negative and 78,786 test positive with IDC. The dataset is available in public domain and you can download it here. You’ll need a minimum of 3.02GB of disk space for this.
Filenames in this dataset look like this:
8863_idx5_x451_y1451_class0
Here, 8863_idx5 is the patient ID, 451 and 1451 are the x- and y- coordinates of the crop, and 0 is the class label (0 denotes absence of IDC).
Prerequisites
You’ll need to install some python packages to be able to run this advanced python project. You can do this with pip-
pip install numpy opencv-python pillow tensorflow keras imutils scikit-learn matplotlib
Steps for Advanced Project in Python – Breast Cancer Classification
1. Download this zip. Unzip it at your preferred location, get there.
Screenshot:
2. Now, inside the inner breast-cancer-classification directory, create directory datasets- inside this, create directory original:
mkdir datasets mkdir datasets\original
3. Download the dataset.
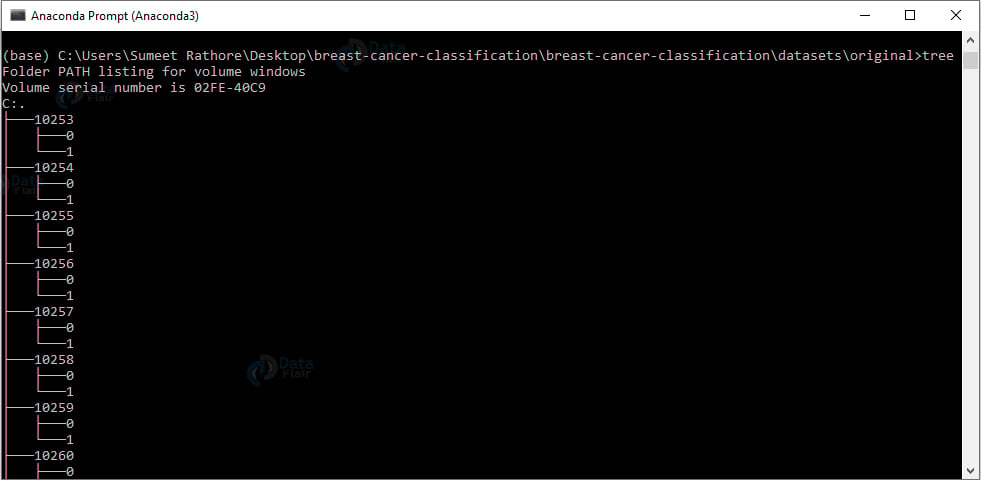
4. Unzip the dataset in the original directory. To observe the structure of this directory, we’ll use the tree command:
cd breast-cancer-classification\breast-cancer-classification\datasets\original tree
Output Screenshot:
We have a directory for each patient ID. And in each such directory, we have the 0 and 1 directories for images with benign and malignant content.
config.py:
This holds some configuration we’ll need for building the dataset and training the model. You’ll find this in the cancernet directory.
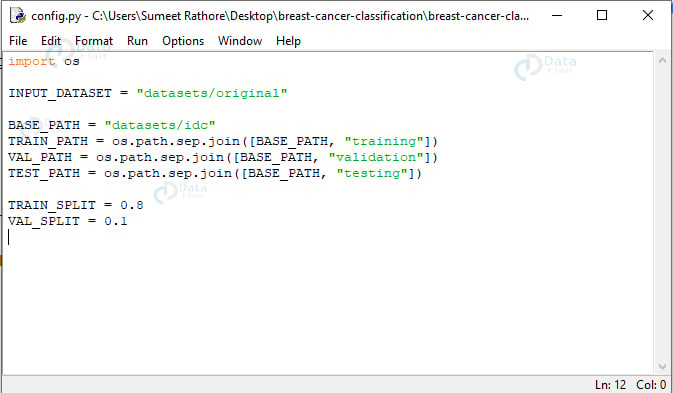
import os INPUT_DATASET = "datasets/original" BASE_PATH = "datasets/idc" TRAIN_PATH = os.path.sep.join([BASE_PATH, "training"]) VAL_PATH = os.path.sep.join([BASE_PATH, "validation"]) TEST_PATH = os.path.sep.join([BASE_PATH, "testing"]) TRAIN_SPLIT = 0.8 VAL_SPLIT = 0.1
Screenshot:
Here, we declare the path to the input dataset (datasets/original), that for the new directory (datasets/idc), and the paths for the training, validation, and testing directories using the base path. We also declare that 80% of the entire dataset will be used for training, and of that, 10% will be used for validation.
build_dataset.py:
This will split our dataset into training, validation, and testing sets in the ratio mentioned above- 80% for training (of that, 10% for validation) and 20% for testing. With the ImageDataGenerator from Keras, we will extract batches of images to avoid making space for the entire dataset in memory at once.
from cancernet import config
from imutils import paths
import random, shutil, os
originalPaths=list(paths.list_images(config.INPUT_DATASET))
random.seed(7)
random.shuffle(originalPaths)
index=int(len(originalPaths)*config.TRAIN_SPLIT)
trainPaths=originalPaths[:index]
testPaths=originalPaths[index:]
index=int(len(trainPaths)*config.VAL_SPLIT)
valPaths=trainPaths[:index]
trainPaths=trainPaths[index:]
datasets=[("training", trainPaths, config.TRAIN_PATH),
("validation", valPaths, config.VAL_PATH),
("testing", testPaths, config.TEST_PATH)
]
for (setType, originalPaths, basePath) in datasets:
print(f'Building {setType} set')
if not os.path.exists(basePath):
print(f'Building directory {base_path}')
os.makedirs(basePath)
for path in originalPaths:
file=path.split(os.path.sep)[-1]
label=file[-5:-4]
labelPath=os.path.sep.join([basePath,label])
if not os.path.exists(labelPath):
print(f'Building directory {labelPath}')
os.makedirs(labelPath)
newPath=os.path.sep.join([labelPath, file])
shutil.copy2(inputPath, newPath)
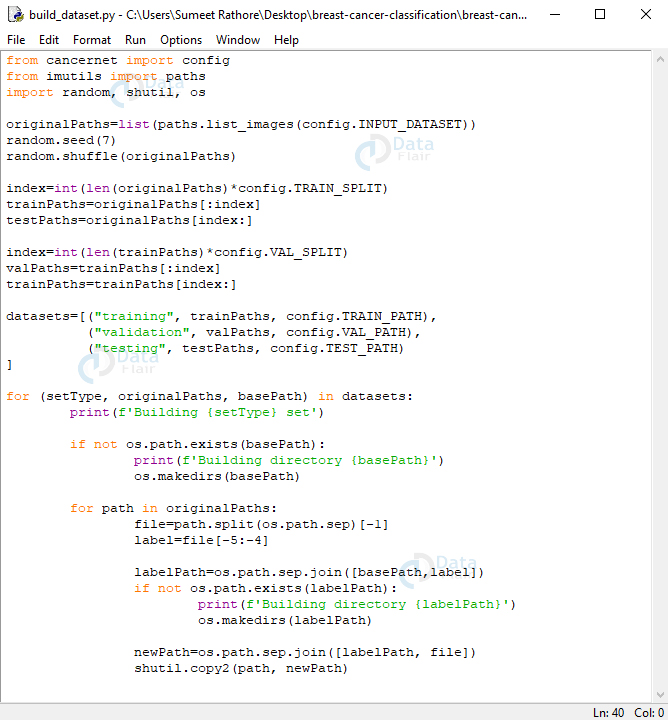
Screenshot:
In this, we’ll import from config, imutils, random, shutil, and os. We’ll build a list of original paths to the images, then shuffle the list. Then, we calculate an index by multiplying the length of this list by 0.8 so we can slice this list to get sublists for the training and testing datasets. Next, we further calculate an index saving 10% of the list for the training dataset for validation and keeping the rest for training itself.
Now, datasets is a list with tuples for information about the training, validation, and testing sets. These hold the paths and the base path for each. For each setType, path, and base path in this list, we’ll print, say, ‘Building testing set’. If the base path does not exist, we’ll create the directory. And for each path in originalPaths, we’ll extract the filename and the class label. We’ll build the path to the label directory(0 or 1)- if it doesn’t exist yet, we’ll explicitly create this directory. Now, we’ll build the path to the resulting image and copy the image here- where it belongs.
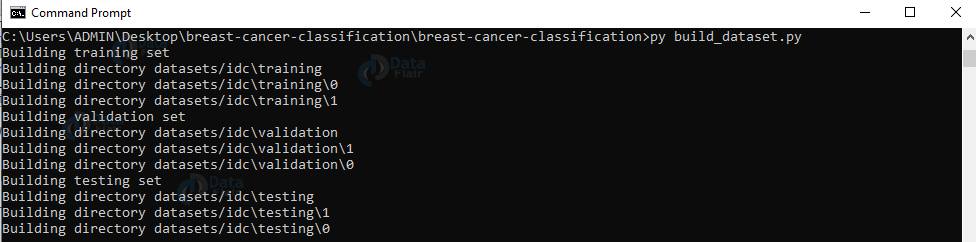
5. Run the script build_dataset.py:
py build_dataset.py
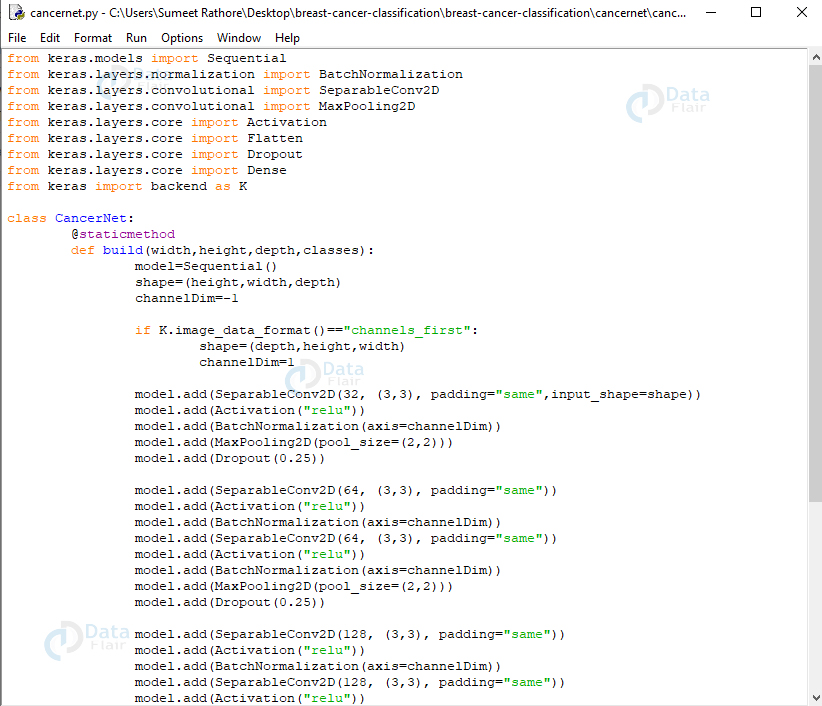
cancernet.py:
The network we’ll build will be a CNN (Convolutional Neural Network) and call it CancerNet. This network performs the following operations:
- Use 3×3 CONV filters
- Stack these filters on top of each other
- Perform max-pooling
- Use depthwise separable convolution (more efficient, takes up less memory)
from keras.models import Sequential
from keras.layers.normalization import BatchNormalization
from keras.layers.convolutional import SeparableConv2D
from keras.layers.convolutional import MaxPooling2D
from keras.layers.core import Activation
from keras.layers.core import Flatten
from keras.layers.core import Dropout
from keras.layers.core import Dense
from keras import backend as K
class CancerNet:
@staticmethod
def build(width,height,depth,classes):
model=Sequential()
shape=(height,width,depth)
channelDim=-1
if K.image_data_format()=="channels_first":
shape=(depth,height,width)
channelDim=1
model.add(SeparableConv2D(32, (3,3), padding="same",input_shape=shape))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(SeparableConv2D(64, (3,3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(SeparableConv2D(64, (3,3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(SeparableConv2D(128, (3,3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(SeparableConv2D(128, (3,3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(SeparableConv2D(128, (3,3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=channelDim))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(classes))
model.add(Activation("softmax"))
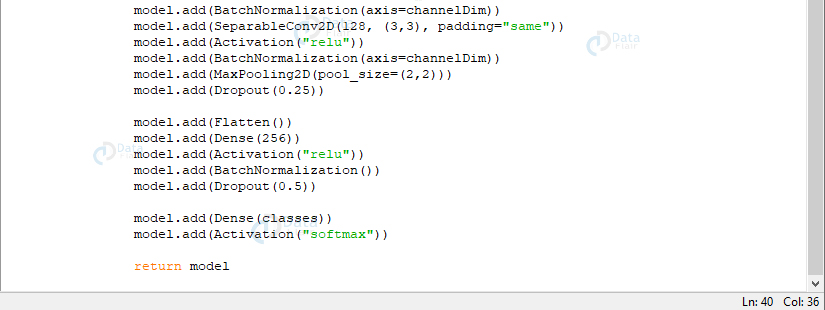
return modelScreenshot:
Screenshot:
We use the Sequential API to build CancerNet and SeparableConv2D to implement depthwise convolutions. The class CancerNet has a static method build that takes four parameters- width and height of the image, its depth (the number of color channels in each image), and the number of classes the network will predict between, which, for us, is 2 (0 and 1).
In this method, we initialize model and shape. When using channels_first, we update the shape and the channel dimension.
Now, we’ll define three DEPTHWISE_CONV => RELU => POOL layers; each with a higher stacking and a greater number of filters. The softmax classifier outputs prediction percentages for each class. In the end, we return the model.
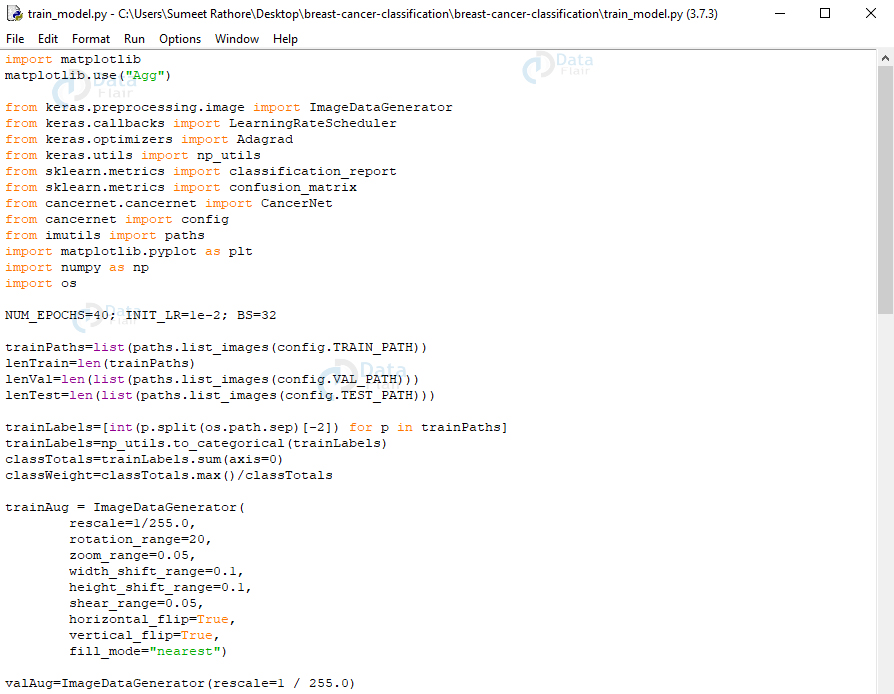
train_model.py:
This trains and evaluates our model. Here, we’ll import from keras, sklearn, cancernet, config, imutils, matplotlib, numpy, and os.
import matplotlib
matplotlib.use("Agg")
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler
from keras.optimizers import Adagrad
from keras.utils import np_utils
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from cancernet.cancernet import CancerNet
from cancernet import config
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import os
NUM_EPOCHS=40; INIT_LR=1e-2; BS=32
trainPaths=list(paths.list_images(config.TRAIN_PATH))
lenTrain=len(trainPaths)
lenVal=len(list(paths.list_images(config.VAL_PATH)))
lenTest=len(list(paths.list_images(config.TEST_PATH)))
trainLabels=[int(p.split(os.path.sep)[-2]) for p in trainPaths]
trainLabels=np_utils.to_categorical(trainLabels)
classTotals=trainLabels.sum(axis=0)
classWeight=classTotals.max()/classTotals
trainAug = ImageDataGenerator(
rescale=1/255.0,
rotation_range=20,
zoom_range=0.05,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.05,
horizontal_flip=True,
vertical_flip=True,
fill_mode="nearest")
valAug=ImageDataGenerator(rescale=1 / 255.0)
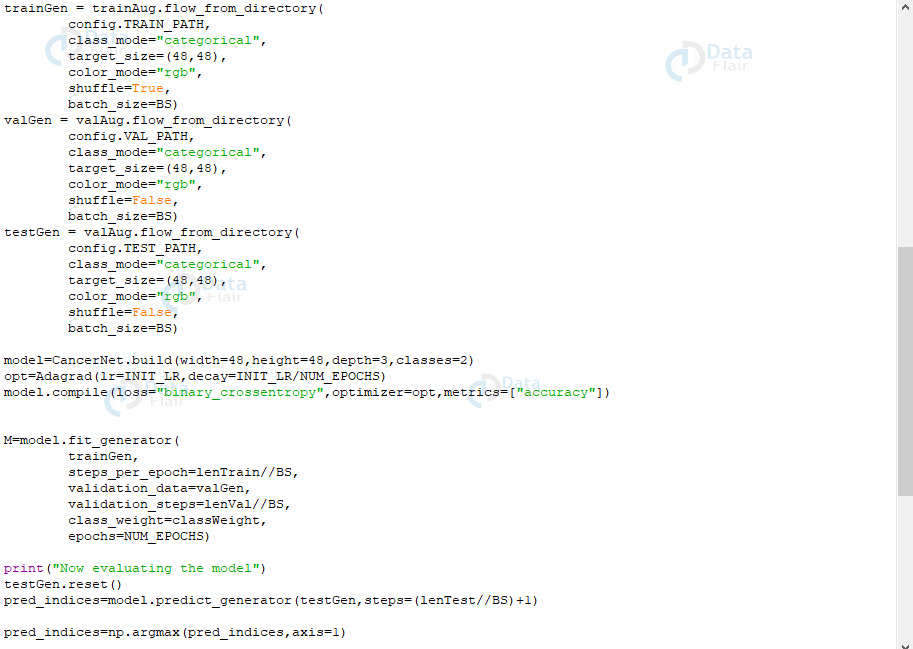
trainGen = trainAug.flow_from_directory(
config.TRAIN_PATH,
class_mode="categorical",
target_size=(48,48),
color_mode="rgb",
shuffle=True,
batch_size=BS)
valGen = valAug.flow_from_directory(
config.VAL_PATH,
class_mode="categorical",
target_size=(48,48),
color_mode="rgb",
shuffle=False,
batch_size=BS)
testGen = valAug.flow_from_directory(
config.TEST_PATH,
class_mode="categorical",
target_size=(48,48),
color_mode="rgb",
shuffle=False,
batch_size=BS)
model=CancerNet.build(width=48,height=48,depth=3,classes=2)
opt=Adagrad(lr=INIT_LR,decay=INIT_LR/NUM_EPOCHS)
model.compile(loss="binary_crossentropy",optimizer=opt,metrics=["accuracy"])
M=model.fit_generator(
trainGen,
steps_per_epoch=lenTrain//BS,
validation_data=valGen,
validation_steps=lenVal//BS,
class_weight=classWeight,
epochs=NUM_EPOCHS)
print("Now evaluating the model")
testGen.reset()
pred_indices=model.predict_generator(testGen,steps=(lenTest//BS)+1)
pred_indices=np.argmax(pred_indices,axis=1)
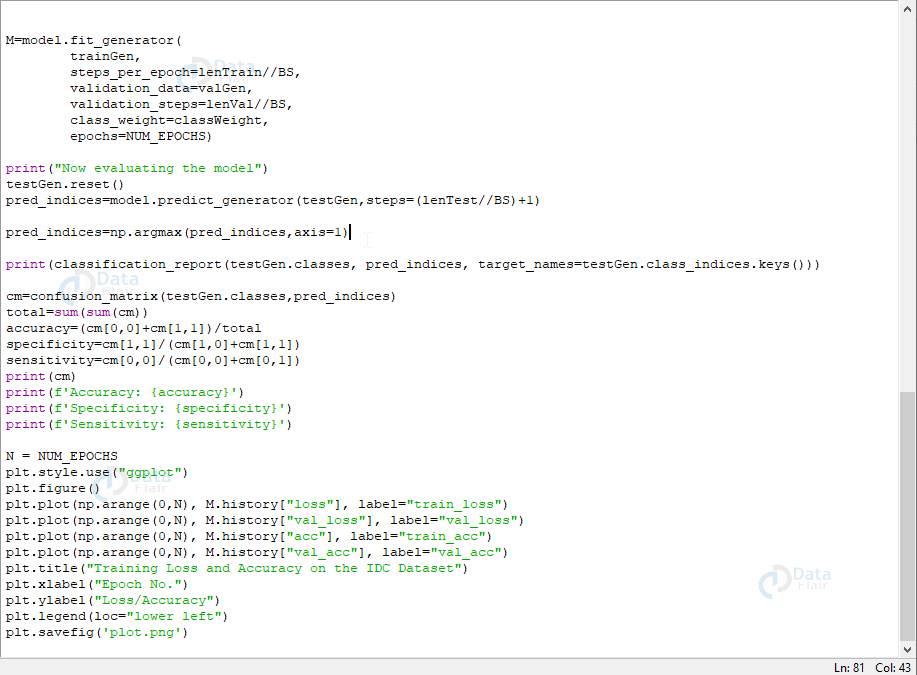
print(classification_report(testGen.classes, pred_indices, target_names=testGen.class_indices.keys()))
cm=confusion_matrix(testGen.classes,pred_indices)
total=sum(sum(cm))
accuracy=(cm[0,0]+cm[1,1])/total
specificity=cm[1,1]/(cm[1,0]+cm[1,1])
sensitivity=cm[0,0]/(cm[0,0]+cm[0,1])
print(cm)
print(f'Accuracy: {accuracy}')
print(f'Specificity: {specificity}')
print(f'Sensitivity: {sensitivity}')
N = NUM_EPOCHS
plt.style.use("ggplot")
plt.figure()
plt.plot(np.arange(0,N), M.history["loss"], label="train_loss")
plt.plot(np.arange(0,N), M.history["val_loss"], label="val_loss")
plt.plot(np.arange(0,N), M.history["acc"], label="train_acc")
plt.plot(np.arange(0,N), M.history["val_acc"], label="val_acc")
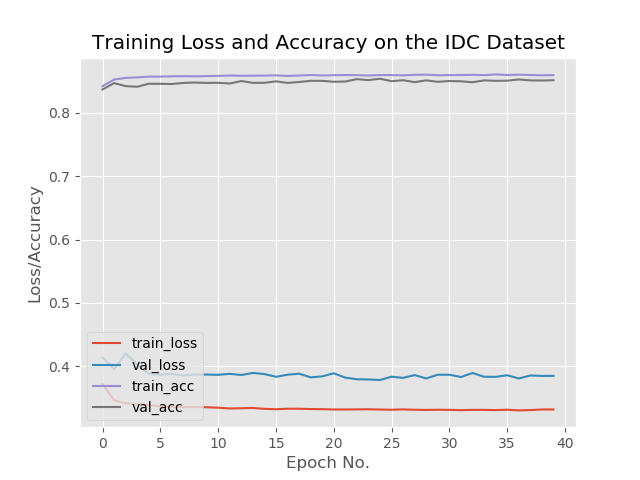
plt.title("Training Loss and Accuracy on the IDC Dataset")
plt.xlabel("Epoch No.")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
plt.savefig('plot.png')Screenshot:
Screenshot:
Screenshot:
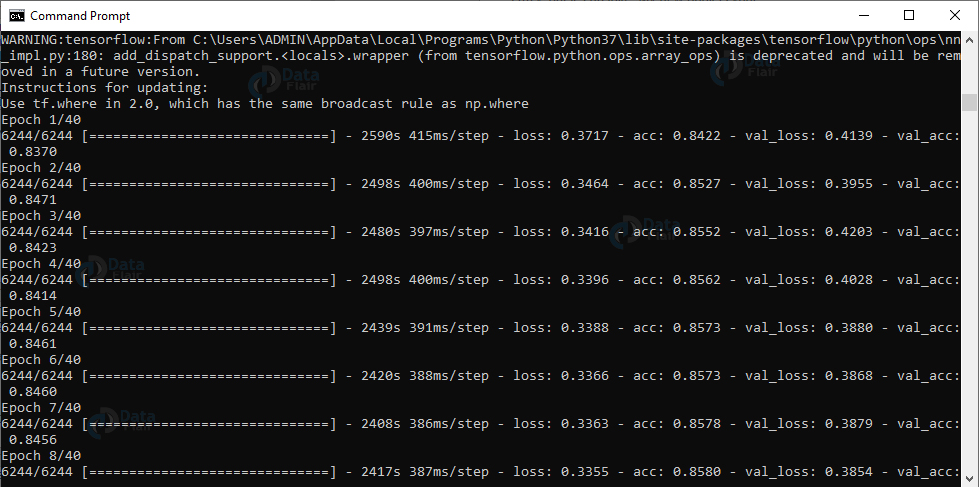
In this script, first, we set initial values for the number of epochs, the learning rate, and the batch size. We’ll get the number of paths in the three directories for training, validation, and testing. Then, we’ll get the class weight for the training data so we can deal with the imbalance.
Now, we initialize the training data augmentation object. This is a process of regularization that helps generalize the model. This is where we slightly modify the training examples to avoid the need for more training data. We’ll initialize the validation and testing data augmentation objects.
We’ll initialize the training, validation, and testing generators so they can generate batches of images of size batch_size. Then, we’ll initialize the model using the Adagrad optimizer and compile it with a binary_crossentropy loss function. Now, to fit the model, we make a call to fit_generator().
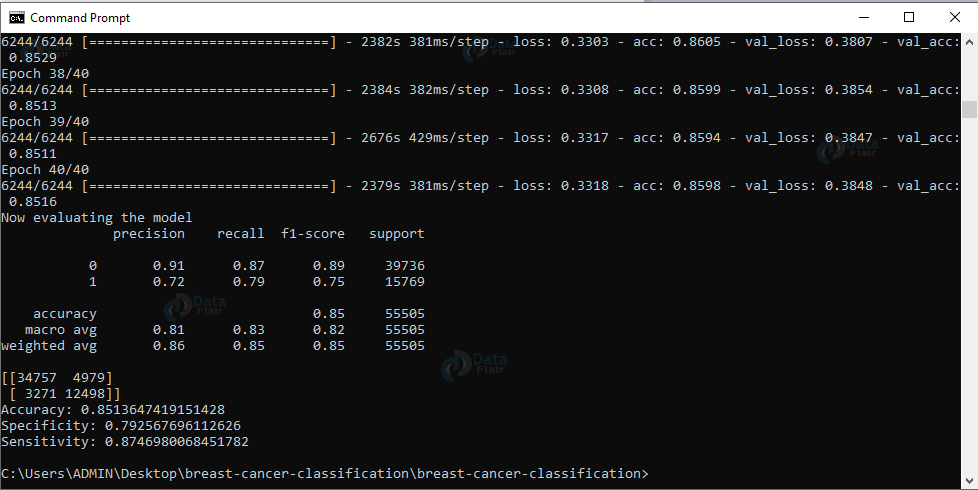
We have successfully trained our model. Now, let’s evaluate the model on our testing data. We’ll reset the generator and make predictions on the data. Then, for images from the testing set, we get the indices of the labels with the corresponding largest predicted probability. And we’ll display a classification report.
Now, we’ll compute the confusion matrix and get the raw accuracy, specificity, and sensitivity, and display all values. Finally, we’ll plot the training loss and accuracy.
Output Screenshot:
Output Screenshot:
Output:
Summary
Breast cancer is one of the most common cancers in women. Detecting it early can save lives. Using machine learning in Python, we can build a classification model to predict if a tumor is malignant (harmful) or benign (safe). This project uses a well-known dataset from the UCI Machine Learning Repository. It includes important features like the size, shape, and texture of cell nuclei from breast mass samples.
In this project in python, we learned to build a breast cancer classifier on the IDC dataset (with histology images for Invasive Ductal Carcinoma) and created the network CancerNet for the same. We used Keras to implement the same. Hope you enjoyed this Python project.
This project is widely used in the healthcare domain to support doctors and improve decision-making. It’s beginner-friendly, yet teaches powerful ML concepts such as classification, data visualization, and model evaluation. By completing this project, you’ll learn to apply machine learning to real-world medical problems and make an impact using data and Python.
Want to become a Data Scientist?
Start learning Python in detail with DataFlair Python Online Training and achieve success.
We work very hard to provide you quality material
Could you take 15 seconds and share your happy experience on Google





Can you send me the dataset if available? It’s not there on kaggle
Hey Nikita,
Here is the dataset of breast cancer classification. You can download it.
Hi Nikita, did you find the dataset to put in the original folder ?
HelloNikita, pls can we connect on twitter?
I have been trying to run the build_dataset.py and all it does is restarts the kernel. does not create folders or split datasets.
I have deduced that the ‘from cancernet import config’ is non-responsive and sends the code to termination.
Please suggest how can I overcome this.
there is an error in this code
from imutils import paths
how to solve it
Hey Mohammed,
Can you specify the error you are receiving? Make sure the package is installed using pip install imutils.
please tell me which python version to use
Can I run this using anaconda and it’s prompt ?
You cannot write the code and run, both in an anaconda prompt. Instead, you have to write in a file with the .py extension and run that file in prompt. You can also run in environments the anaconda provides like Jupyter Notebook, Spyder, etc. Hoping that this answered your query.
which code to run after the build_dataset.py,
please state the steps till the end
We first declare all the paths in config.py. Then, we divide the dataset and split it into batches in build_dataset.py. After this, we build the CNN model in the file cancernet.py. Finally, we train and test the model by writing the code in train_model.py. Hoping this explanation gave a clear picture of the model.
Which python version to use,
Because i am getting error in tensorflow and more
It is best to have the Python 3 versions to run the algorithms. It is also in demand and gives advatanges over the older versions.
thank you very much, it worked perfectly, even though i run it on CPU, and it took some quite time. I think the model is not saved on the disk, so if i want to run the model once again for other unseed pictures, i have to run it again to save it first, right?
Yes, if you want to work on different pictures or data set then you have to follow the comlete procedure again.
Please can’t find data to put in the original folder (they are not avalable in kaggle)
Do you mean to get the dataset for training and testing? If yes, then you can download from the drive https://drive.google.com/file/d/1nEkiRNIdYUSi0Eyci19KceLJjObGB25m/view . In this article, we have used this dataset. If you want to work on a different dataset, then you can download a similar dataset by searching in Kaggle. Hoping that your question is answered.
Please can’t find data to put in the original folder (they are not avalable in kaggle)
Could you please tell me the approximate run time? I tried to run the build_dataset.py file and it’s just stuck at building training set 1. It won’t go beyond that no matter how much time I wait. It just kept on running for about 3.30 hrs.
Very good work.Well done. I want to build a Deep Learning model, using a Genetic Algorithm to optimize the hyper parameters. My dataset is going to be from customs transactions. Can you please assist with implementation guide? Thank you.
same issue as Neethu
did anyone manage to fix it?
nether mind i just had to wait and use py instead of python
Hi buddy can you help me with how to execute these programs please if i have to make this for my college
You can follow the similar work done, that is saving code in different python files and running them in the anaconda prompt. You can also use Jupyter Notebook and write code in different cells. In addition, you can also add information using Markdown. Or else, you can use other environments as well.
Hi, How can we visualize the result for a testing pack? Let say I need to test a new patient mammogram. Thank you
You can store the new information in a dataset and let the trainGen get worked on this dataset. Hope this helps.
Dear author, please help me to fix this error
if class_weight:
ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all()
Dear sir, did you found any solution to this error?
Did you find any solution for this error
help me .
ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all()
Hi did you find any solution for this error
Did u find any solution for this error
Hi there, I had the same issue, here is how I fixed it.
It seems like this was written when tf used model.fit_generator() instead of model.fit().
model.fit() takes in a dictionary for the class_weight parameter. As it is in the code above it is a numpy array. To fix this you need to create a dictionary that contains the weights and the corresponding label. Something like
classWeightDict={0: classWeight[0], 1:classWeight[1]} then pass that to model.fit(… ,… ,class_weights=classWeightDict)
Hi there, I had the same issue, here is how I fixed it.
It seems like this was written when tf used model.fit_generator() instead of model.fit().
model.fit() takes in a dictionary for the class_weight parameter. As it is in the code above it is a numpy array. To fix this you need to create a dictionary that contains the weights and the corresponding label. Something like
classWeightDict={0: classWeight[0], 1:classWeight[1]} then pass that to model.fit(… ,… ,class_weights=classWeightDict)
I am getting error in model.fit_generator(epochs=NUM_EPOCHS)
the error is value error
Can anyone help
Please share the link to dataset.
I am not able to find it anywhere else.
https://www.kaggle.com/paultimothymooney/breast-histopathology-images/. The dataset is available on this link.
The link to the data set is https://drive.google.com/file/d/1nEkiRNIdYUSi0Eyci19KceLJjObGB25m/view . You can download this zip file and extract the files.
Should we build a cancernet or is it built already because when we run the program the error says ” no module named ‘cancernet’ “
Hello Dear,
How can I plot the confusion matrix for this project (please provide the code or command line that i should add it to print it)
Thanks in advance
Hi,
In the given code, the variable cm is holding the confusion matrix. To plot this you can use the seaborn library, which can be installed by using the “”pip install seaborn”” or “”conda install seaborn””. Then write the following code import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
df_cm = pd.DataFrame(cm, range(2), range(2))
sn.heatmap(df_cm, annot=True)
plt.show() Hope this helps!”
I’m getting an error while installing the packages. It says ” Could not find a version that satisfies the requirement tensorflow”.
Tensorflow works for all Python 3 versions. You can actually check your version by writing the command “python –version” in terminal. You can update the Python by using the command “apt update”. Hope I could solve your query.
I am unable to unzip the data from “https://www.kaggle.com/paultimothymooney/breast-histopathology-images/” after downloading the zip. How should I use the dataset.
Can someone help me out please
After downloading, you can right-click on the zip and select Extract All.. option. This will extract all the files in the location of your choice. These files are the datasets we use. Hope this solves the issue.
if class_weight:
ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all()
Can someone help me out with this error
I’m getting an error with such as PIL.UnidentifiedImageError: cannot identify image file <_io.BytesIO object at 0x000002535483CD60. I can't seem to resolve the issue.
Actually, this error generally shows up in training datasets. You can solve this by attempting to open all images using the command im = Image.open() and validate by calling im.load() all of your images. Using this, you can record the filenames that fail. Another option is to include the code in the try and catch block. Hope this helps you solve the problem.
It is a very interesing project to be worked on but I need some help in it. I am facing some issues.
Why in “cancernet.py” do we repeat the functions SeparableConv2D(), Activation() and BatchNormalization() 6 times? Once with SeparableConv2D(32,…), twice with SeparableConv2D(64,…) and three times with SeparableConv2D(128,…).
This is because we have six different convolution layers with each having its characteristics. Having different layers helps in improving the model. Hope I could answer your question.
Sir finally after completion of project we will find the accuracy of CNN on classification .
But how can we give input images for classification after project
You can follow a similar procedure. First, you build the dataset. Then instead of splitting it into train and test, we give it as input to the model built. And get the accuracy.
Anyone have DDSM breast cancer dataset. If you have please with me.
Can u send me the full project if possible..?
I am having this error:
cannot import name ‘BatchNormalization’ from ‘keras.layers.normalization’
Could you please guide me on this? Thanks
when i build dateset . only tarining dateste had been build . validation and test had not been build . IDE has took a long time and not thing can you help my where is my rong ??